Gene transcription is facilitated by RNA polymerase enzyme complexes that collaborate with transcription factors, repressors, chromatin remodelers, and other cellular factors. RNA Polymerase III (RNAPIII) mainly transcribes short DNA fragments called tDNAs, that code for transfer-RNAs (tRNAs). In repressive conditions, tDNA transcription is repressed by the well-characterized protein Maf1. A new study by Van Breugel et al., recently published in Molecular Cell, identified Fpt1p (YKR011C) as an additional regulator of RNAPIII in S. cerevisiae.
By using Epi-Decoder, a technique based on synthetic genetic array (SGA), chromatin immunoprecipitation and DNA-barcode sequencing, the local chromatin-proteome of a single tDNA was decoded in active and repressive conditions. The authors found major reprogramming of the core RNAPIII transcription machinery and other known chromatin-binding proteins. Surprisingly, they found the protein Ykr011c to be enriched in the tDNA chromatin-proteome, especially under repressive conditions, prompting the authors to rename the gene FPT1 (Factor in the Proteome of tDNAs number 1).
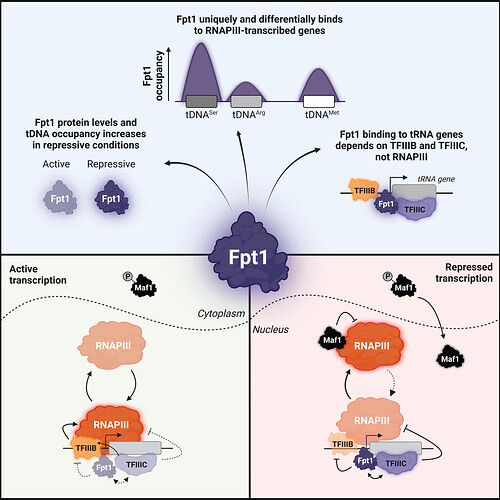
Following up on the Epi-Decoder finding, genome-wide sequencing methods such as ChIP-seq and ChIP-exo revealed that Fpt1p uniquely binds RNAPIII-regulated genes. Using the anchor away system to conditionally deplete core RNAPIII transcription factors from the nucleus, Fpt1 binding to tRNA genes was found to require both TFIIIB and TFIIIC but not RNAPIII or ongoing transcription. tRNA genes have been described to differentially respond to repressive signals but gene-specific regulatory mechanisms have largely remained elusive. Looking at Fpt1p, Van Breugel et al. found a correlation between tDNA responsiveness to repressive signals and Fpt1p occupancy, suggesting a negative regulatory role for Fpt1p. Substantiating these results, FPT1 knockout strains showed increased occupancy of RNAPIII and TFIIIB at tRNA genes, while TFIIIC occupancy decreased. These outcomes point towards a role for Fpt1p in promoting eviction of RNAPIII upon repressive signals.
In summary, taking advantage of multiple yeast genetic approaches, Van Breugel et al. found that the previously uncharacterized protein Fpt1 is a bona fide RNAPIII regulator in S. cerevisiae. Their research emphasizes the importance of not overlooking uncharacterized proteins, as they may possess alternative regulatory roles that could change our views on fundamental cellular processes.
Text and image provided by Marlize van Breugel, MSc.