A 2022 study has used a modified yeast strain “ABC16-Green Monster” with fewer export pumps (i.e. more susceptible to drugs) to exert powerful selection pressure on yeast cells. By exposing this strain to multiple libraries of chemical compounds with potential use as antifungal agents, Ottilie et al. in a 2022 issue of Communications Biology identified an evolved set of 25 genes with frequent mutations.
In comparing the set of mutations to those frequently found upon extended growth without selection, they noted that intergenic mutations were comparatively rare, presumably due to the heavy selection pressure for functional resistance. For another confirmation of the screen’s effectiveness in isolating useful variants, i.e. not passenger mutations, they introduced 61 of the altered alleles back into the unevolved strain by CRISPR/Cas9 integration and verified that 45 variants across 37 genes restored resistance.
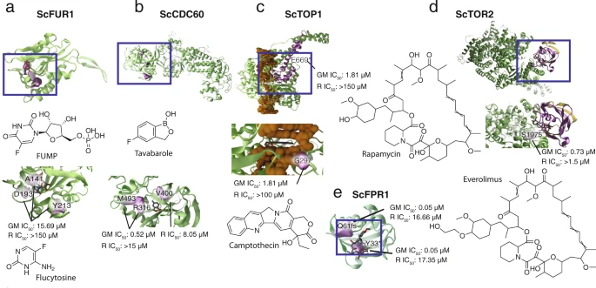
Looking more closely at the mechanisms of action for the observed resistances, they noted mutations clustered in the active sites of target molecules for which the target was previously known. For example, the antifungal drug tavaborole (a type of benzoxaborole) led to identification of four active-site mutations in Cdc60p (the yeast leucyl-tRNA synthetase) that are predicted to interfere with binding to the drug. In a related example, the chemotherapy drug camptothecin inhibits topoisomerase and they isolated two mutations in TOP1 that would be expected to fall in the binding pocket.
Their isolation of mutations in TOR2 and FPR1 (both associated with TOR signaling) in strains with evolved resistance to rapamycin reproduce the findings of a related study looking at rapamycin resistance in yeast. In fact, both studies identified mutations in the same S1975 residue of Tor2p. In other interesting findings, they isolated mutations leading to yeast resistance to drugs used typically against soil-transmitted helminths (worms), trypanosomatid parasites, and malarial Plasmodium. Each mutant variant in yeast affords insight into how these pathogens might likewise evolve resistance or could be alternately targeted.
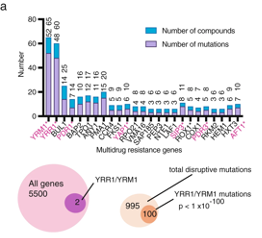
The set of identified mutant genes was highly enriched for transcription factors and among these were two - YRR1 and its paralog YRM1 - that together mediated resistance for nearly 25% of the compounds tested. These two zinc transcription factors had previously been shown to activate genes involved in multidrug resistance. In this study, they were mutated 100 times in screens against 19 diverse chemical compounds. Interestingly, deletion of these genes does not confer resistance and the sum of the data suggests the screen identified gain-of-function alleles. In support of this idea, integration of the L611F allele of YRR1 into a susceptible strain by CRISPR/Cas9 reconstructed resistance to a suite of compounds. The researchers hypothesize that modified proteins lead to constitutive activation of genes aiding resistance.
In sum, this study demonstrates the awesome power of yeast genetics (#APOYG) for revealing insight into the molecular underpinnings of medical chemistry. Studies like this one provide not only data but intriguing clues about where to look next—most of which will be easy to test in yeast.