Gene transcription — the elaborate process that our cells use to read genetic information stored in DNA – was long thought to be turned on only when certain regulatory factors traveled to specific DNA sequences. In a recent study published in Genes & Development, Mittal et al., 2022, discovered that a subset of genes has their transcription regulatory factors and cofactors already in place, but in a latent state. With the appropriate signals, these “poised” genes become highly active.
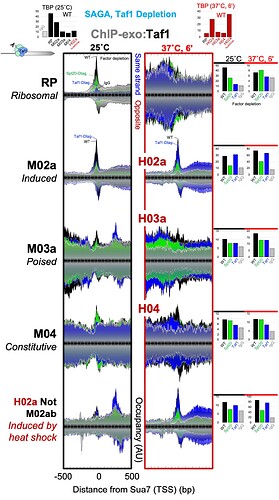
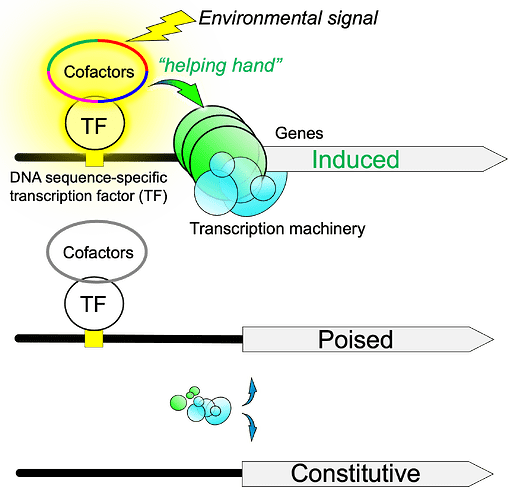
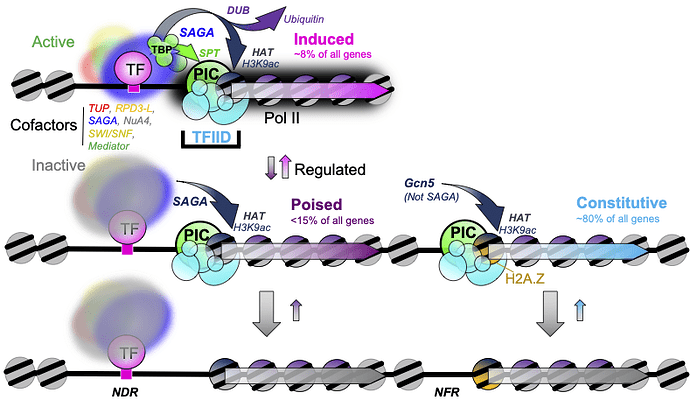
The authors calculated the DNA-bound preinitiation complex/TBP-associated factor (PIC/TAF) ratio at all yeast genes and identified two major classes. The first, and largest, group provides basic “housekeeping” functions and are usually “on” at very low levels all the time (i.e., “constitutive”). The second class, the “inducible” genes, has a whole entourage of “poised” proteins assembled nearby which provides a guiding hand to transcription machinery when triggered by environmental signals, resulting in high levels of “induced” transcription.
Using CRISPR-Cas9 mediated protein depletion/degradation and gene knockout techniques, Mittal et al. removed parts of the SAGA and TFIID cofactors to systematically examine the role they play in regulating the above gene classes. They discovered that the constitutive class largely depended on TFIID, whereas the inducible genes required both SAGA and TFIID, suggesting an integrated pathway of PIC assembly at inducible genes. If true, such a possibility would be different from previous models where the two cofactors were thought to engage in somewhat distinct mechanisms of PIC assembly.
The authors further examined the above results and were surprised to find that at the inducible promoters, SAGA stabilized Taf1p (and thus TFIID) which helped in a rapid and robust transcription initiation of these genes upon acute environmental changes.
In addition, Mittal et al. also removed gene-specific transcription factors (TFs) to address how these factors recruited TBP upon sensing changes in environmental signals. They showed that TFs such as Hsf1p and associated cofactors were already bound to gene promoters prior to induction, instead of traveling to cognate sites upon induction. These cofactors, thus, lend a helping hand in recruiting TBP and the associated machinery at the induced genes.
Lastly, Mittal et al. removed Gcn5p and DUB subunits of SAGA to examine the effect on histone acetylation and ubiquitylation, respectively. They showed that global histone acetylation and ubiquitylation levels require active Gcn5 and the deubiquitination (DUB) activities, suggesting a SAGA independent moiety contributing towards maintaining total cellular acetylated and ubiquitylated histone pools.
Image credits: Image 1 – Mittal et al., 2022; Image 2 – B. Fanklin Pugh, Cornell University; Image 3 – Mittal et al., 2022.
Contributed by Chitvan Mittal.